During their research, funded in part by the Spanish Cystic Fibrosis Foundation and the Obra Social “La Caixa”, the scientists were looking at Pseudomonas aeruginosa bacterium strain PAO1, the reference strain in the majority of clinical and research laboratories, in their quest to find new strategies to combat bacterial infections.
Above: PAO1 bacteria infecting zebrafish with the mutation in the promoter of nrdD (upper panel, less green) and with a strain that has corrected the mutation in the nrdD promoter (lower panel, much more green).
One enzyme that is essential for cell division in PAO1 is ribonucleotide reductase RNR, which supplies the monomers required for DNA synthesis and repair. P. aeruginosa is one of the few microorganisms that encodes three different RNR classes in its genome, enabling it to grow and adapt to diverse environmental conditions, including during infection – which is why it has become so ubiquitous in microbiology laboratories.
The researchers found that a lack of RNR activity leads to cell elongation in PAO1, which indicates growth. Not only that, but RNR gene expression during anaerobiosis – processes occurring without oxygen – differs between clinical P. aeruginosa strains and those used in the laboratory, with class III RNR being much more highly expressed in clinical P. aeruginosa isolates compared to the lab strain.
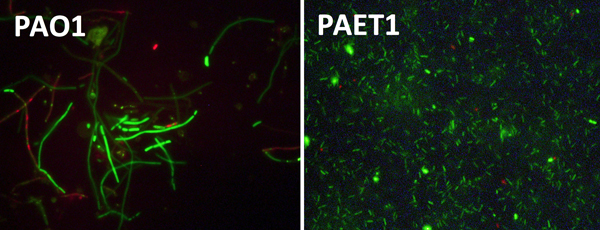
Right: Bacterial cells of PAO1 (left) and clinical isolates (right) grown anaerobically. PAO1 shows clear growth defects with elongated cells compared to the clinical isolate.
“It appears that PAO1 has adapted to laboratory conditions, a bit like when our bodies become resistant to antibiotics,” says Eduard. “During infection, clinical P. aeruginosa acts under anaerobic conditions, but the P. aeruginosa lab strain PAO1 can’t grow anaerobically. We found, though, that a single mutation in the class III RNR promoter region of the PAO1 strain disrupts its anaerobic transcription, decreasing its virulence. This means PAO1 is inappropriate for anaerobic or infection studies, and nor is it good for when you’re searching for new antimicrobial compounds that involve anaerobic conditions.”
The discovery was made during Eduard’s work developing new strategies to fight bacterial infections and possible therapeutic targets, which is strongly linked to cystic fibrosis. This disease causes the formation and accumulation of thick mucus in the lungs, intestines, pancreas and liver, and patients suffer respiratory failure due to chronic bacterial colonization – biofilms – in the lungs. Eduard’s research is taking new approaches to overcome this problem, such as developing testing systems that closely resemble what is found in a patient’s lungs, as well as introducing several nanomedicine strategies to combat these infections.
At the end of 2016 the group received a generous boost over three years thanks to a new funding initiative from Obra Social “La Caixa”, which made the current co-operation triangle of researchers, clinicians and the patient’s association possible.
—
Source article: Anna Crespo, Joan Gavaldà, Esther Julián & Eduard Torrents (2017). “A single point mutation in class III ribonucleotide reductase promoter renders Pseudomonas aeruginosa PAO1 inefficient for anaerobic growth and infection”. Scientific Reports, 7, Article number: 13350





