Group: Cellular and molecular mechanobiology
Group leader: Pere Roca Cusachs (proca@ibecbarcelona.eu)
A major hallmark in bioengineering is the ability to engineer cells to carry out specific tasks, which can then be applied to problems in regenerative medicine, cancer, and many other pathological scenarios. To be useful, engineered cells must respond to their environment. Specifically, our lab has recently shown that a fundamental mechanism by which cells sense their environment is by force transmission to the cell nucleus, which changes its shape and leads to the nuclear import of transcriptional regulators [1]. This leads to the exciting hypothesis that nuclear tension and shape control cell function, and that this understanding can be harnessed to design cell systems with a biological output that can be controlled by force.
The objective of this project will be to engineer cells that respond to applied forces, by triggering a response of choice. This will be implemented by designing and testing molecular constructs that specifically enter the nucleus with force, and activate specific transcriptional programs. Depending on the case, this response may include cell migration, proliferation, or differentiation, among others.
The expertise acquired will include biomechanical tools, molecular cell biology techniques, and computational modelling, and will also involve collaborations with leading international groups. The student will be integrated in a highly interdisciplinary group composed of physicists, engineers, biologists, and chemists.
Elosegui-Artola E, Andreu I, Beedle AEM, Lezamiz A, Uroz M, Kosmalska AJ, Oria R, Kechagia JZ, Rico-Lastres P, Le Roux AL, et al.: Force triggers YAP nuclear entry by regulating transport across nuclear pores. Cell 2017:In press.
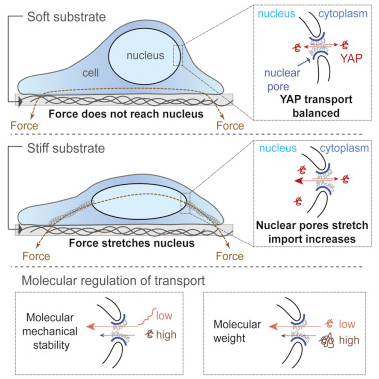
Proposed mechanism of regulation of control of cell function by force-dependent nuclear import of the transcriptional regulator YAP (Elosegui-Artola et al., Cell 2017).

