Former group leader: Antonio Juárez
About
1. Structure and function of bacterial proteins that modulate virulence expression

Protein–protein and protein–DNA interactions play key roles in the ability of virulent bacteria to adapt to the host environment and cause disease. A group of proteins is currently the focus of our research: nucleoid-associated proteins (NAPs) that contribute to DNA architecture and modulate gene expression. We are interested in unravelling the role played by two of these proteins – Hha and H-NS – in the regulation of virulence and of plasmid transfer. Escherichia coli pathotypes such as enteroaggregative E. coli are the subject of our research. Owing to their key modulatory functions, these proteins are interesting targets to combat bacterial infections.
2. Bacterial plasmids and their role in transmission of multidrug resistance markers
A main concern with bacterial infections is the selection of isolates that are resistant to several antimicrobial drugs. The transmission of the ability of bacterial cells of simultaneously resist several antimicrobial drugs is accomplished, in many instances, by plasmids. These genetic elements can be transmitted from one cell to another, and modify the phenotype of the recipient cell. We have recently shown that multidrug resistance plasmids in Salmonella require specific plasmid proteins to be stably maintained in this microorganism. These proteins could be considered as targets to combat multidrug resistance.
3. Application of nanotools of bacterial biotechnology
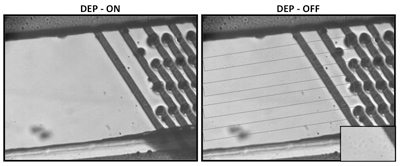
3.1. Dielectrophoresis. We have previously shown that dielectrophoresis can be a valuable tool for bacterial cell sorting and characterization. We are currently using different chip designs (2D and 3D carbon electrodes) to: a) study the effect of electric fields on bacterial cell physiology; b) combine DEP with other molecular protocols for detection and identification of different types of cells. Recent results have shown that DEP chips can be used to increase PCR detection of yeast cells.
3.2. Atomic force microscopy (AFM). Conventional AFM approaches have been shown to be powerful techniques for characterizing both biomaterials and biomolecules. In a joint project with the Nanoscale Bioelectrical Characterization group (page 76), we intend to use electrical- AFM to characterize the bacterial cell envelope. We also plan to use this approach to analyze the structural and physiological properties of bacterial living cells.
Projects
National projects
| Regulación de la virulencia bacteriana por proteínas que reconocen conformaciones locales del ADN | REGVIRBAC | Antonio Juárez |
| INTERMODS Interconexiones de Módulos plasmídicos y los Genomas de Bacterias Patógenas | MINECO-CSIC | Antonio Juárez (managed by UB) |
Privately funded projects
| MEJORAVE1 Mejora sanitaria y de productos cármicos de ave | Industrial project with Mevet, S.A – CZ Veterinaria, S.A | Antonio Juárez |

